
For a single pixel, one common definition is the lowest intensity that can be identified with a 99% confidence (example, greater than or equal to three standard deviations of baseline noise.) The limit of detection is the lowest intensity that can be confidently identified within a background. This method provides a rough comparison of sensitivity, but if only intensity is evaluated and not measurements of signal to background or signal to noise, the system's limit of detection can be misestimated. Band intensity is often used to compare imaging systems. When evaluating candidate imaging systems for purchase, many researchers will take images of the same blot using identical (or near identical) settings. After all, the whole western blotting experiment can take upwards of two days so a few extra minutes during image acquisition is a small price to pay to see a low-expressing protein. It is important to consider the instrument with the lower ultimate limit of detection, as long as it takes a reasonable amount of time to acquire that image. While quick and easy to perform side-by-side, this method compares time to results, rather than the ultimate sensitivity of the imagers. Often, two imagers are evaluated by imaging the same blot with identical imaging times and comparing the resulting band intensities. In western blotting, sensitivity of imaging instruments is often discussed using two somewhat differing definitions. This requires optimization of all the upstream steps of the western blotting workflow as well as an imaging system that has the highest sensitivity possible. I'd be happy to have answers in the form of hotkeys, macros, or entirely different solutions.One of the most common challenges for a western blotting experiment is the detection of low-abundance protein targets. Is there a way to perform the measurement step for all the rectangles in the lane at once? Second, when doing the measuring step, currently I must click on each rectangle individually, then hit m, then manually select the next one, press m again, etc., like so: Is there an easy way to move all the rectangles downward the same number of pixels at the same time? In many programs, something like Shift + Click or Command + Click would allow one to select multiple rectangles, for instance. csv (I am about to restart at step 4 for lane 2). Here, I've just completed the process and exported the results to. My immediate goal for this post is really to speed these few steps up (below), unless there is an easy generalized solution. However, there are a few key steps that if those could be streamlined, the process would become very rapid. for lane 2 and repeat above steps starting from 4.Īutomating this entire process would be best, but strikes me as a bit more involved than I have time for at present for a number of reasons. 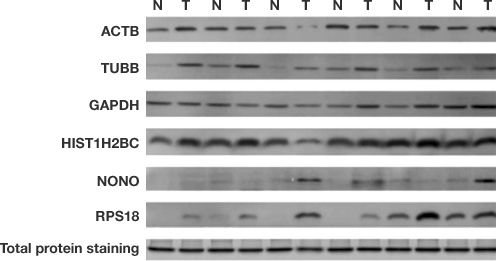
Finally, export all the measurements to a. Select each rectangle, and press `m` each time. Click on the first rectangle (from Step 5.). Repeat until every band in the first lane is selected. 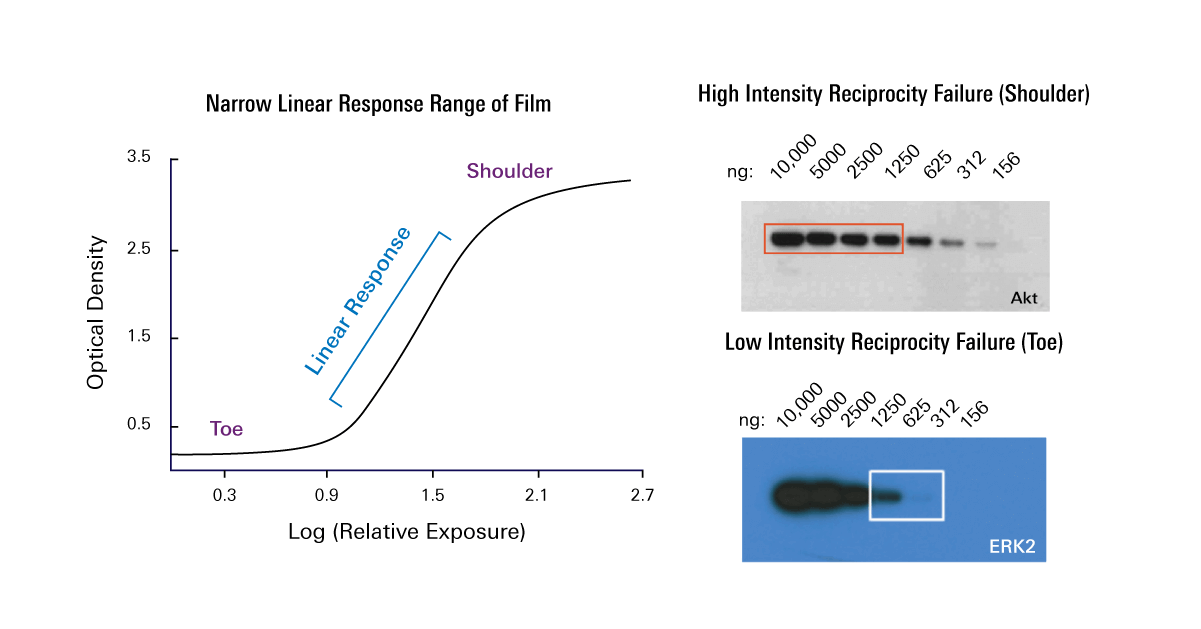
Move rectangle to second band -> Select next lane.ħ. Go to Analysis -> Gels -> Select first lane.Ħ. Draw a first rectangle around the first band in lane 1.ĥ. Select Edit, then `Invert` the image (such that white -> black black -> white)Ĥ. My basic procedures follows NIH recommendations for use of ImageJ. However, if there are relevant features in the standalone versions, please let me know as I would be willing to switch versions. For reproducibility, I will be using the web-based version for this post. Using ImageJ for the first time, in this case to quantify Western Blot data.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
